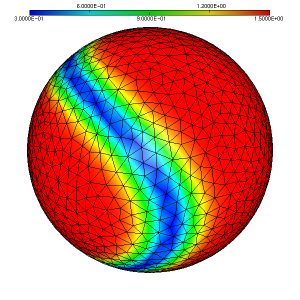
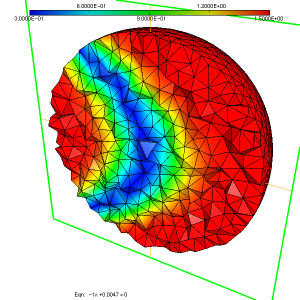
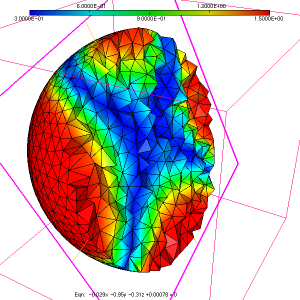
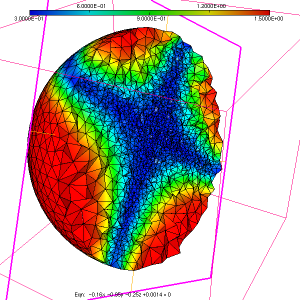
As a simple test case to illustrate the usage of the ParMmg application, we consider the initial isotropic mesh of a sphere named sphere.mesh, with the associated sizemap sphere.sol displayed in figure 1, figure 2 and figure 3.
We run the application in parallel on 8 processors.
mpirun -np 8 parmmg_O3 sphere -out sphere-out.mesh -mesh-size 200000 -niter 6 -nlayers 3 -hgradreq 3.0
- the -out option allows to choose the output file name.
- the -mesh-size option controls the size (in elements) of the mesh parts to be sequentially remeshed on each process (not needed in this small example).
- the -niter option controls the number of parallel remeshing-repartitioning iterations.
- the –nlayers option controls by how many mesh edges a parallel interface is displaced to repartition the mesh. A value of 2 or 3 is recommended (default is 2; a too high value can worsen load balancing).
- the -hgradreq option controls the gradation near parallel interfaces (a value between 2.0 and 4.0 is suggested; default is 2.3).
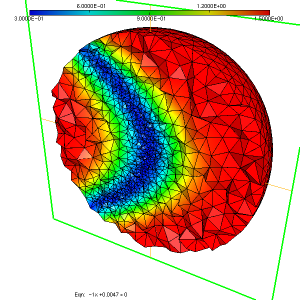
We obtain a mesh named sphere-out.mesh and the associated sizemap (see figure 4, figure 5). This sizemap is saved in the sphere-out.sol file.
Remark:
- The ParMmg application accepts most of the command line options accepted by Mmg. More help can be found by typing
parmmg_O3 -h
- Adaptation on boundary surfaces is currently not supported (i.e. option -nosurf is applied by default).
- This is a small test case intended to illustrate how to use the application from command line (Parmmg is targeted to handle large distributed meshes through its library). No speedup with respect to mmg3d is expected in this case, because the computational cost of iteratively remeshing and repartitioning overcomes the benefits of the parallel execution for such a small input mesh.